Publication: New methodology for characterization of cell-free genetic networks enables forward engineering
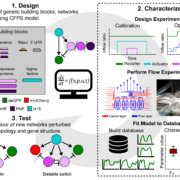
Forward engineering of (cell-free) genetic networks is one of the main goals in the field of synthetic biology. However, despite the vast library of available building blocks for the assembly of these networks, identifying the reaction kinetics to make these building blocks modular for forward engineering remains elusive. In our latest work we established an automated pipeline to produce high-quality time-resolved data through a combination of optimal experimental design, microfluidics and non-linear model identification. We apply this pipeline and progress through a design, characterize test cycle and demonstrate the modularity and predictability of the characterized building blocks in new network configurations. The methodology described in this paper has the potential to enable forward engineering of cell-free genetic networks.
You can find the article here: https://www.nature.com/articles/s41467-022-31306-3
Congratulations Bob, Roel, Wilhelm and collaborators!